
Long DNA palindromes are difficult to directly analyze using standard molecular genetics methods. It will be useful for studying human cancers and other diseases associated with palindromes. The GAP-Seq approach underscores the importance of developing new tools for identifying and characterizing palindromes, and provides a new strategy to systematically assess palindromes in genomes. Because this protocol eliminates many of the false positives that plague earlier techniques, we have improved palindrome identification. Using MCF-7 breast cancer cell line as the proof-of-principle analysis, we have identified 35 palindrome candidates and physically characterized the top 5 candidates and their junctions. It also indicates the location of the novel junction, which can then be recovered. Unlike earlier protocols, it does not involve restriction enzymatic digestion prior to DNA snap-back thereby preserving longer DNA sequences. We developed a new protocol to identify palindromes that couples the S1 nuclease treated Cot0 DNA (GAPF) with high-throughput sequencing (GAP-Seq). Because of their role in genomic instability and gene amplification in some human cancers, it is important to develop systematic approaches to detect and characterize DNA palindromes. The ability to form these hairpins results in genome instability, difficulties in maintaining clones in Escherichia coli and major problems for most DNA sequencing approaches. Here, for example, the single difference in the sequences can be eliminated (red for blue or vice versa).Closely spaced long inverted repeats, also known as DNA palindromes, can undergo intrastrand annealing to form DNA hairpins. Repairs can then be made (probably by the mechanism of homologous recombination). All that is needed is to form a loop so that the two sequences line up side-by-side. This orientation and redundancy may help ensure that a deleterious mutation in one copy of the set can be repaired using the information in another copy of that set. GTGTT AAGGGTACCCAACACCCTC 5' - 3' GAGGGTGTTGGGTACCCT AAACAC. CACAA TTCCCATGGGTTGTGGGAG 3' - 5' CTCCCACAACCCATGGGA TTTGTG. (The dashes represent the thousands of base pairs that separate adjacent palindromes.)ĥ'. The human Y chromosome contains 7 sets of genes - each set containing from 2 to 6 nearly-identical genes - oriented back-to-back or head-to-head that is, they are inverted repeats like the portion shown here. Inverted repeats at either end of retroviral gene sequences aid in inserting the DNA copy into the DNA of the host.The DNA of many transposons is flanked by inverted repeats such as this one:ĥ' GGCCAGTCACAATGG.~400 nt.CCATTGTGACTGGCC 3'ģ' CCGGTCAGTGTTACC.~400 nt.GGTAACACTGACCGG 5'.This graphic shows the "recognition helix" to which the CAP protein (a homodimer) binds in the lac operon of E. Transcription factors are often dimers of identical proteins (" homodimers") so it is not surprising that each member of the pair needs to "see" the same DNA sequence in the same orientation. The DNA sequence shown above is that of the glucocorticoid response element where n represents any nucleotide. The DNA to which transcription factors bind. 
In these cases, two different segments of the double helix read the same but in opposite directions. The graphic shows the palindromic sequences "seen" by five restriction enzymes (named in blue) commonly used in recombinant DNA work. This type of palindrome serves as the target for most restriction enzymes. 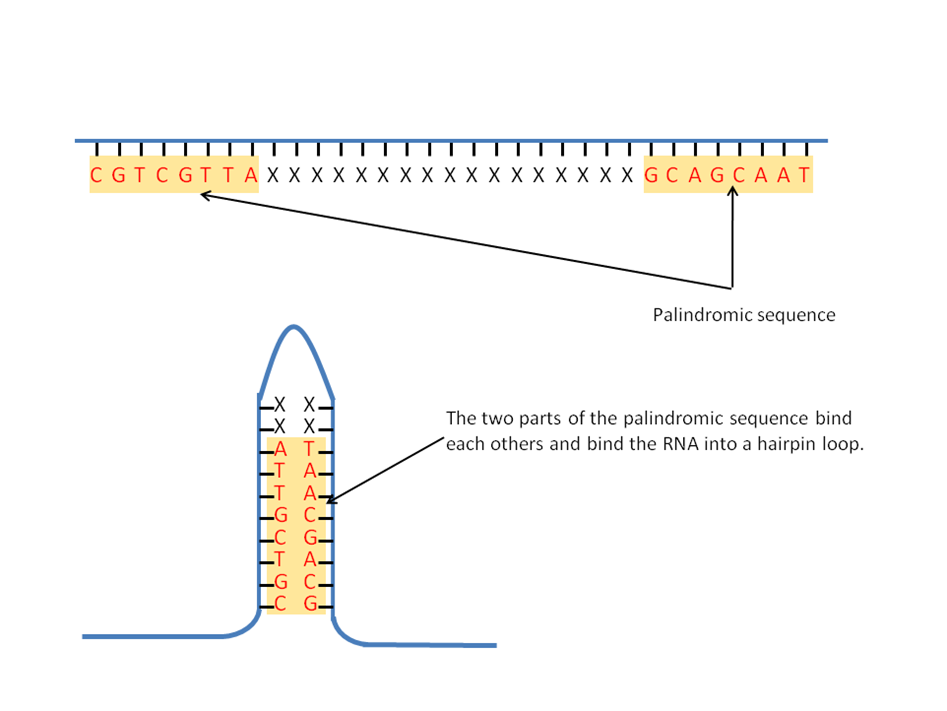
Palindromes that occur on opposite strands of the same section of DNA helix. " able was I ere I saw elba" is a palindrome.ġ. A palindrome is a sequence of letters and/or words, that reads the same forwards and backwards.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
